Algorithm — NNDescent
Consider n data points X. We want to find k nearest neighbors of every point x in X. Here, we are given some function d(x1,x2) that computes some distance between the two points. Often times, this is Euclidean distance, but any arbitrary function will work too. Let’s say the time complexity of the distance function is O(d).
The brute force approach would be to go through ever pair within X and record the pairwise distance, and select minimum k neighbors for every point x. The complexity would be O(n*(n*d + n*log(k))), since for each point x, we need to compute pairwise distance with all other points and do partial-sort to select nearest k points. This results in n^2 term, which is infeasible for any n that is reasonably large, say a tens of thousands or so.
As with any algorithm, we can significantly reduce the time complexity if we sacrifice the accuracy. That is, if we are willing to accept k neighbors that may not be true nearest neighbors but close. We would measure this accuracy in terms of recall, which is the ratio of true neighbors in the k points recalled by the algorithm.
One of such algorithm is called nearest neighbor descent or NNDescent, introduced by this paper. The technique relies on a simple idea: a neighbor’s neighbor is likely a neighbor. Here is how we generate nearest k-neighbors for each point using NNDescent algorithm:
- For each point
xinX, randomly pickkpoints to be thek-neighbors. - In each iteration, we compute distance between
xand all of its neighbors’ neighbors. We update itsk-neighbors. We do this for every pointxinX. - We repeat the iterations until there is no further update of neighbors for the entire data
X.
This is a very simple yet effective algorithm. Below shows a time evolution of # updates and recall for each iteration, ran with n = 10,000, k = 10, and dim = 10 and Euclidean distance.
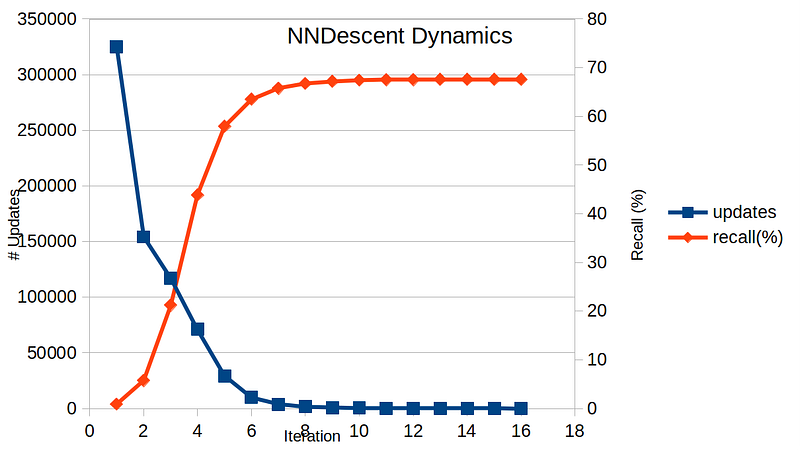
Here, # updates refers to the total count of updated neighbors in one iteration. The stopping condition is when # updates is zero. In this particular example, it took 16 iterations to converge. Looking at the recall values, we see that the result is not optimal — the recall is capped at below 70%. How much of time did we save with this approximate technique?
The time complexity of NNDescent would be O(m * n * (k^2 * d + k^2 * log(k))), where m is the # iterations. Given that typically m, k << n, we can see that this technique reduces complexity quite a lot. The authors claim O(n^1.14) found empirically on various dataset.
So, here are my experimental results by varying k, dim, and n to get a rough idea of sensitivity on various parameters. The default parameters are n = 10,000, k = 10, and dim = 10.
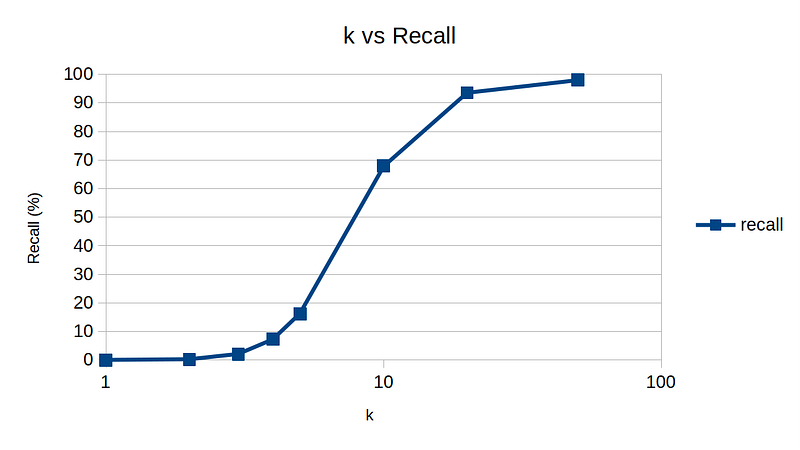
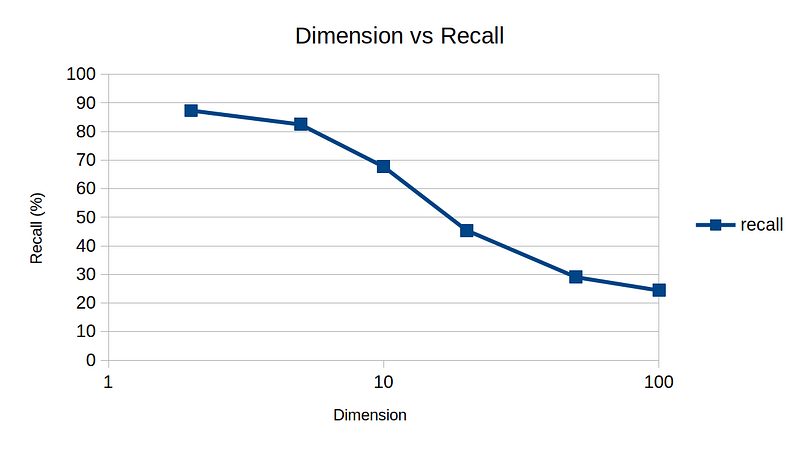
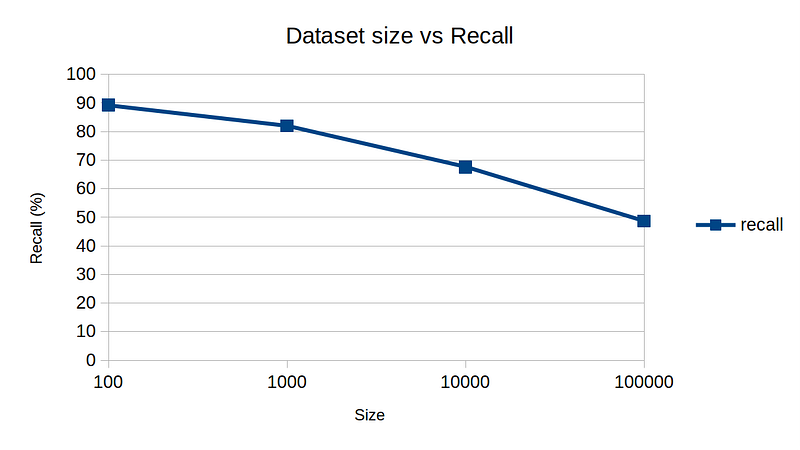
One interesting note is dimension vs recall plot. For high-dimensional data, NNDescent technique is not as effective, again demonstrating the curse of dimensionality.